This is just a dumping ground for commands I perform all the time so that I have somewhere to look them up!
Convert tophat bam file into a sorted sam file for htseq-count
#if used tophat should be already sorted
samtools view -h tophat_sorted.bam > tophat_sorted.sam
#if need to sort before conver (output tophat_sorted.bam)
samtools sort tophat.bam tophat_sorted
Bash
wc -l seq_file.fas #count lines
grep -c '>' seq_file.fas #count seqs in fasta file
ls -l #permissions and file size
ls -lh #above with human format for sizes
chmod a+wx #add read write permissions to all
chmod 755 #read and execute for user/world
chmod -R 755 #change all files in folder
tail -n 400 #last 400 lines
tail -n +400 #from 400 to end
history | grep "command" #search history for commands
#sed prints to screen, redirect or us -i for inplace
#print a specific line (100) using sed -n just shows that line
sed -n -e '100d' test.txt
#delete a specfic line (line 1000) from a file and backup original
sed -i.bak -e '1000d' file.txt
#delete lines
sed -i.bak -e '1d,10d
#removes line 10 to 20 INCLUSIVE ie 11 lines
sed -i -e '10,20d' test.txt
#remove line 10, lines 15-20, line 100
sed -i -e '10d;15,20d;100d' test.txt
#convert fastq to fasta file!(effecient)[p is print]
sed -n '1~4 s/^@/>/p;2~4p' seq.fq > seq.fas
#get the md5 checksum of a folder
find FOLDER/ -type f -exec md5sum {} \; | sort -k 34 | md5sum > md5.txt
Htseq-count -s flag turn off stranded union is default
htseq-count -s no -m union tophat_sorted.sam > sample_counts.txt
Vim
v #visual
p #paste
# find word at hash
yy #yank word or line
d #delete line
wq #save and close
ma #mark a, use 'a' to goto this section
mz # mark z as above
{} #select paragraph
{d} #cut block</pre>
Python specific
pip install --upgrade packagename
#virtual env
virtualenv venv
source venv/bin/activate
deactivate venv
print "%f"%(np.mean(seq_length))
#prints 1019.662143
print "%.2f"%(np.mean(seq_length))
#prints 1019.66
print "%.1f"%(np.mean(seq_length))
#prints 1019.7 note rounding up
print "%d"%(np.mean(seq_length))
#prints 1019
#use with open('') as for automatic file closing
with open('infile.txt','r') as f:
f.read()
Git
WordPress (see text for tags)
use the following tags in sqaure_brackets code language="css" and end with code in sqr bracktets
import sys
test = sys.argv[1]
for name in test:
print name
plot(xmir205,ycbx1,'go',fig_fn205(xmir205'),'-k')
plot(xmir205,ycbx1,'go',fig_fn205(xmir205),'-k')
R
browseVignettes()
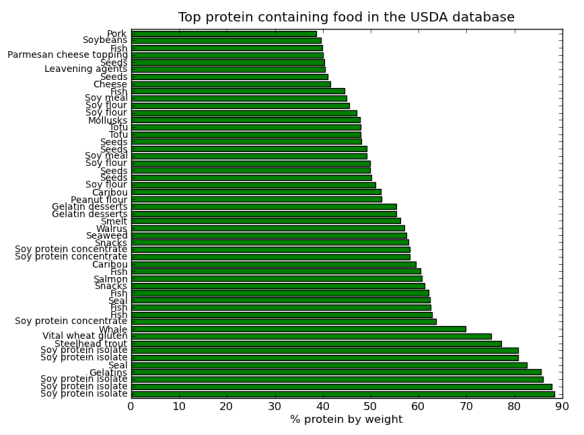
Matplotlib
Change the font family.
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 22}
matplotlib.rc('font', **font)