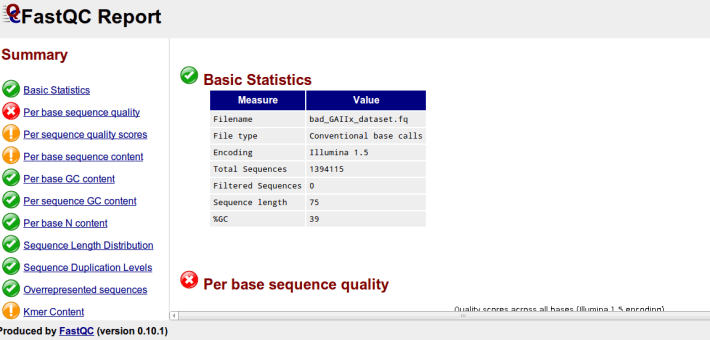
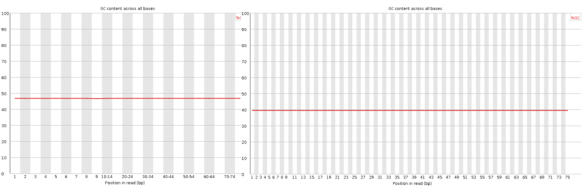
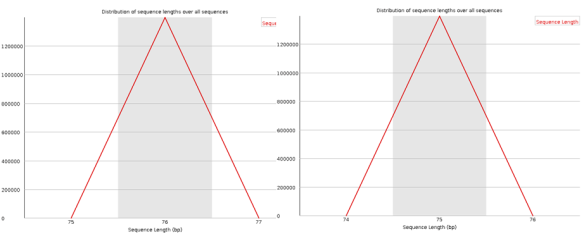
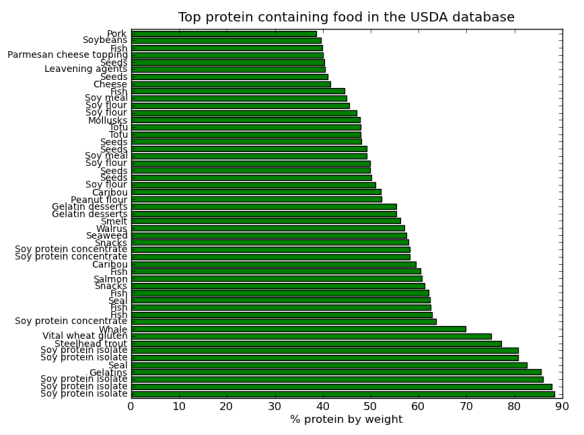
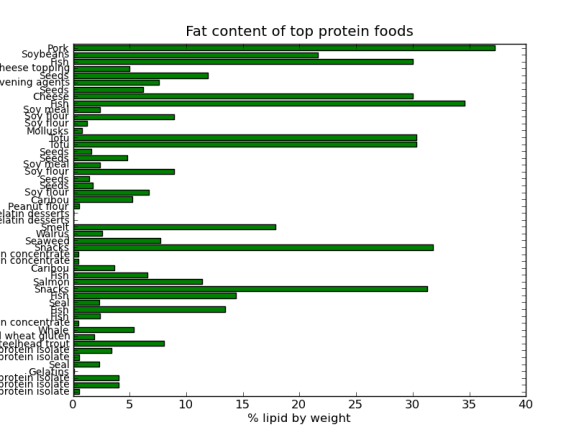
I’ve posted before on how much I love plotting in python using pandas, matplotlib, and ipython. But I thought I would do a blog post that would contain some important ‘bookmarks’ on how to do some essential plotting tasks. Just as a simple example of how pandas could be used in a biological sense, how about we make a dataframe from a blast table outputted from NCBI BLAST. The example data is just a blast search using transcripts as a qry with output option 8, which creates a tabular output that is nice for parsing using scripts. Note, if you want to use my table go for it, its here as an tab-delimited Excel spreadsheet, for this example I just copied it into a text document called ‘blast_output.txt’ – I can’t upload txt files at wordpress)-:)
1) Basic table manipulations and very simple excel-like maths functions.
Normally I probably would pre-process the data using a simple python script, but here I’ll do it all in pandas.
from pandas import Series,DataFrame
import matplotlib.pyplot as plt
import pandas as pd
blast_data = pd.read_table('blast_output.txt',header = 0)
blast_data = DataFrame(blast_data)
blast_data.head()
# QUERY NUMHITS HITDESC \
#0 Sequence_7116 1 ref|XP_004521769.1| PREDICTED: M-phase phospho...
#1 Sequence_7117 1 ref|XP_002059188.1| GJ16167 [Drosophila virili...
#2 Sequence_7118 1 ref|XP_002008154.1| GI11977 [Drosophila mojave...
#3 Sequence_7119 1 ref|XP_001955873.1| GF24906 [Drosophila ananas...
#4 Sequence_7120 1 ref|XP_004520750.1| PREDICTED: proteoglycan 4-...
#
# NUMHSP EVAL BITSCORE PERCENTIDENT PERCENTCONSERV QUERYCOV \
#0 1 2.000000e-44 158 0.702290 0.793893 0.511719
#1 1 1.000000e-175 502 0.766667 0.886667 0.853081
#2 3 1.000000e-46 132 0.680412 0.762887 0.126928
#3 1 2.000000e-62 211 0.452830 0.577358 0.716599
#4 2 0.000000e+00 734 0.514886 0.643608 0.577610
# HITCOV
#0 0.796178
#1 0.974026
#2 0.572414
#3 0.933824
#4 0.650243
So now we have the blast table stored as a pandas data frame, it might be a good time to show how to splice and dice this frame so that it shows information that we are interested in. Firstly, column and index headings are easy to access using (in this example) “blast_data.columns” and “blast_data.index”. To grab a column is simple:
blast_data['QUERY'] ##output## 0 Sequence_7116 1 Sequence_7117 2 Sequence_7118 3 Sequence_7119 4 Sequence_7120 5 Sequence_7121 ......
To slice section of the table we would use “blast_data.ix”. As this is a two demensional table sometimes its a little tricky to workout how “ix” works. But its quite simple really, its just the two dimensions separated by a comma, with the row range first and columns second, remembering that as this is python counting starts at 0! So for some easy examples, lets see how to get the first line and all rows (1), first 10 rows and 3 columns (2), first 10 rows and last 2 columns (3). As you can see its just normal python list splicing but in 2 dimensions.
#first line and all rows (1)
blast_data.ix[0:0,:] #index 0 to 0 ie first row and ":" columns ie all of them
###output###
QUERY NUMHITS HITDESC \
0 Sequence_7116 1 ref|XP_004521769.1| PREDICTED: M-phase phospho...
NUMHSP EVAL BITSCORE PERCENTIDENT PERCENTCONSERV QUERYCOV \
0 1 2.000000e-44 158 0.70229 0.793893 0.511719
HITCOV
0 0.796178
#first 10 rows and 3 columns (2)
blast_data.ix[0:10,0:3] #standard python slice
###output####
QUERY NUMHITS HITDESC
0 Sequence_7116 1 ref|XP_004521769.1| PREDICTED: M-phase phospho...
1 Sequence_7117 1 ref|XP_002059188.1| GJ16167 [Drosophila virili...
2 Sequence_7118 1 ref|XP_002008154.1| GI11977 [Drosophila mojave...
3 Sequence_7119 1 ref|XP_001955873.1| GF24906 [Drosophila ananas...
4 Sequence_7120 1 ref|XP_004520750.1| PREDICTED: proteoglycan 4-...
5 Sequence_7121 1 ref|XP_004537614.1| PREDICTED: protein suppres...
6 Sequence_7122 1 ref|XP_002073069.1| GK13353 [Drosophila willis...
7 Sequence_7123 1 ref|XP_001981946.1| GG11315 [Drosophila erecta...
8 Sequence_7124 1 ref|XP_002052580.1| GJ17621 [Drosophila virili...
9 Sequence_7125 1 ref|XP_002007909.1| GI13202 [Drosophila mojave...
10 Sequence_7126 1 ref|XP_004520332.1| PREDICTED: nucleolar prote...
#first 10 lines and last 2 rows (3)
blast_data.ix[0:10,-2:] #standard python again, '-2' is 2 from end
###output###
QUERYCOV HITCOV
0 0.511719 0.796178
1 0.853081 0.974026
2 0.126928 0.572414
3 0.716599 0.933824
4 0.577610 0.650243
5 0.312649 0.376129
6 0.615196 0.996016
7 0.934222 1.000000
8 0.840226 1.000000
9 0.157009 0.342857
10 0.837484 0.960961
The great thing about the pandas dataframe is you can do excel like maths work using the table, for example firstly lets just look at E-values < 1E-100.
good_hits = blast_data[blast_data['EVAL']<1e-100]
len(good_hits)
#output 576
good_hits.head()
###output####
QUERY NUMHITS HITDESC \
1 Sequence_7117 1 ref|XP_002059188.1| GJ16167 [Drosophila virili...
4 Sequence_7120 1 ref|XP_004520750.1| PREDICTED: proteoglycan 4-...
5 Sequence_7121 1 ref|XP_004537614.1| PREDICTED: protein suppres...
6 Sequence_7122 1 ref|XP_002073069.1| GK13353 [Drosophila willis...
7 Sequence_7123 1 ref|XP_001981946.1| GG11315 [Drosophila erecta...
NUMHSP EVAL BITSCORE PERCENTIDENT PERCENTCONSERV QUERYCOV \
1 1 1.000000e-175 502 0.766667 0.886667 0.853081
4 2 0.000000e+00 734 0.514886 0.643608 0.577610
5 2 1.000000e-108 390 0.438655 0.605042 0.312649
6 1 1.000000e-112 342 0.648221 0.833992 0.615196
7 1 0.000000e+00 1845 0.513501 0.672667 0.934222
HITCOV
1 0.974026
4 0.650243
5 0.376129
6 0.996016
7 1.000000
We can even collect the best hits by sorting by the EVAL column.
best_hits.sort('EVAL',inplace=True)
best_hits.head()
###output##
QUERY NUMHITS \
501 Sequence_7617 1
400 Sequence_7516 1
754 Sequence_7870 1
398 Sequence_7514 1
755 Sequence_7871 1
HITDESC NUMHSP EVAL \
501 ref|XP_004525475.1| PREDICTED: kelch domain-co... 1 0
400 ref|XP_001362065.2| GA19724 [Drosophila pseudo... 1 0
754 ref|XP_004524178.1| PREDICTED: mitogen-activat... 1 0
398 ref|XP_004523028.1| PREDICTED: sphingosine kin... 1 0
755 ref|XP_004536256.1| PREDICTED: V-type proton A... 1 0
BITSCORE PERCENTIDENT PERCENTCONSERV QUERYCOV HITCOV
501 691 74 0.863248 0.537193 0.902724
400 842 61 0.744382 0.904181 0.973574
754 2147 73 0.837891 0.934356 0.965003
398 636 53 0.680445 0.797900 0.993399
755 897 84 0.897143 0.379302 0.949367
Some *very* simple math might be to convert the percent identity into a percentage value
good_hits['PERCENTIDENT'] = good_hits['PERCENTIDENT']*100 good_hits['PERCENTIDENT'] = good_hits['PERCENTIDENT'].astype(int)
Of course you could do a lot more complex excel type maths than that, but hopefully you get the idea!
2) Histograms – everyone loves a good histogram.
For this histogram how about we collect information from a blast search and parse out the species, so that we can look at histogram of ‘species richness’ in the best BLAST hits. In this format column 3 contains a hit descriptor that also lists the species name in square brackets (ie [Nasonia vitripennis]). So I want to parse out that information, collect all the totals, and plot the most frequent 10 (for example). We really only need the species part, so we discard the genus information to avoid problems with all those contaminating Drosophila genomes in the database (-;
Now we are really interested in the column “HITDESC”. What we want is just the frequency distribution of the species in the square brackets, all the other info is not required for a histogram.
desc = blast_data['HITDESC'] desc ###output### 0 ref|XP_004521769.1| PREDICTED: M-phase phospho... 1 ref|XP_002059188.1| GJ16167 [Drosophila virili... 2 ref|XP_002008154.1| GI11977 [Drosophila mojave... 3 ref|XP_001955873.1| GF24906 [Drosophila ananas. ......
Although names are descriptions are incomplete this is just a display issue, the data is still there. First we will write a quick filtering function to grab the species name, then use list comprehension to make a collection of names, finally, we will use another function to count each occurrence of the names in the list.
def species(desc):
#a function to pull out the species name from [species genus]
line = line.split(' [')[1] #get species genus
line = line.split(' ')[0] #trim off genus part
return line
def counter(seq):
"""make a freq dict with species as key"""
seq_dict = {}
for n in seq:
if n in seq_dict:
seq_dict[n] += 1
else:
seq_dict[n] = 1
return seq_dict
desc_list = [species(des) for des in desc] #make a list of names
###output###
['Ceratitis',
'Ceratitis',
.......
'Ceratitis',
'Oikopleura']
desc_dict = counter(desc_list)
#lambda magic to get a sorted list from an unsorted dictionary
most_freq8 = sorted(spec_dict,key=lambda x:spec_dict[x],reverse=True)[:8]
#if you wanted to check this works
#chk = spec_dict.values()
#chk.sort()
#chk[-8:]#show last ten values, crossref these against diction usin#g key as most freq species from most_freq8
Nearly there, you’ll probably notice our most_freq8 list is no good for a histogram because we have lost our frequency information. We can get it back using some functional programing magic (BTW for an excellent run through of functional programing in python check out the great how-to-here here).
most_freq_dict = dict(((spec,spec_dict[spec]) for spec in most_freq8))
most_freq_dict
###output###
{'Acyrthosiphon': 7,
'Aedes': 8,
'Anopheles': 6,
'Ceratitis': 501,
'Drosophila': 388,
'Glossina': 7,
'Musca': 7,
'Sarcophaga': 8}
Briefly, a dictionary can be created from tuple of key value pairs, so all we do here is use list comprehension to combine the “spec_dict” frequency counts with the keys from the most frequent list to create a new dictionary. As it turns out, dictionaries can be converted directly to pandas dataframes, however, this will create a frame with all the species as column headings. We could flip the axes using DataFrame.transpose fucntion, but that is messy. Instead we will use list comprehension again to reformat the data (Yes I know I probably could have done this a few steps ago to avoid all this data mushing).
hist_data = DataFrame([[key,most_freq_dict[key]] for key in most_freq_dict])
hist_data
###output###
0 1
0 Musca 7
1 Aedes 8
2 Drosophila 388
3 Anopheles 6
4 Ceratitis 501
5 Acyrthosiphon 7
6 Sarcophaga 8
7 Glossina 7
#need the index to be the species to get them to display
Great, sort of! Really, we want the species names to be the index so they are displayed in the histogram, we could just re-index from here, but lets re-make the table directly specifying that we want the index to be the key from the dictionary
hist_data = DataFrame([most_freq_dict[key] for key in most_freq_dict],columns = ['Frequency'],index=most_freq_dict.keys())
hist_data #we use the same dictionary(most_freq_dict) for columns and index, so this will work!
###Output###
Frequency
Musca 7
Aedes 8
Drosophila 388
Anopheles 6
Ceratitis 501
Acyrthosiphon 7
Sarcophaga 8
Glossina 7
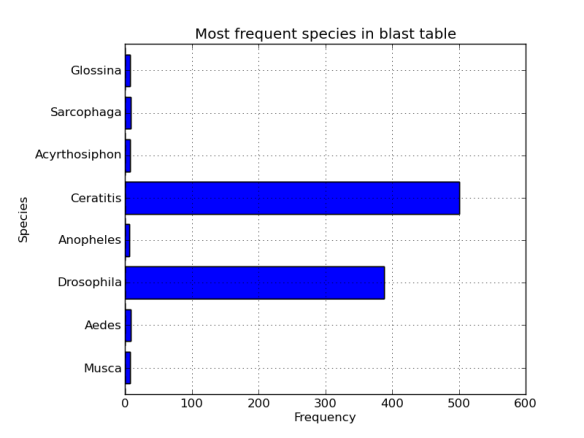
We can make a histogram directly from out dataframe, sweet.
fig = plt.figure()
axes = fig.add_subplot(1,1,1)
axes.set_ylabel("Species")
axes.set_xlabel("Frequency")
axes.set_title("Bla bla")
hist_data.hist(kind='barh',ax=axes,legend=False)
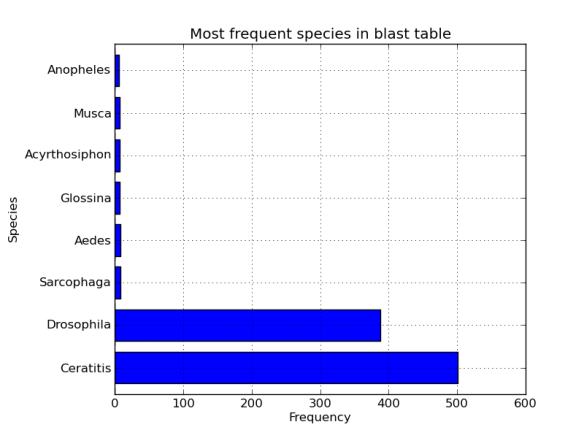
Finally, we really want the data_frame sorted by size to make the histogram look a little nicer.
hist_data = hist_data.sort('Frequency',ascending=False) #not in place
axes = fig.add_subplot(1,1,1)
axes.set_ylabel("Species")
axes.set_xlabel("Frequency")
axes.set_title("Bla bla")
hist_data.hist(kind='barh',ax=axes,legend=False)